Replies: 6 comments 1 reply
-
|
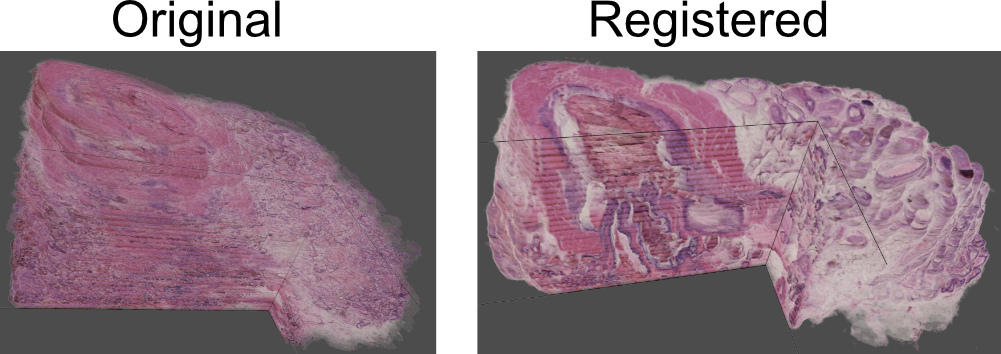
Hi @NichD, this sounds like a very cool project! It is true that valis was originally intended for 2D registration, but I recently tested valis to see how it performs with 3d histology reconstruction. The dataset came from Kartasalo et al 2018, and involved aligning 260 serially sliced images. The authors provided landmarks to asses alignment quality, and for the prostate sample the error was 11um, which is an improvement over the reference result presented in the paper (15um). Here's a figure I was putting together for the website that shows the before and after: So, the registration remains at its core a series of pairwise alignments, but it seems to work well for 3d registration. I did make a few small adjustments to the code to improve the quality of 3d registration, which will be in the next update (1.0.0rc13). However, in the meantime you should be able to get similar results if you set Best, |
Beta Was this translation helpful? Give feedback.
-
|
Hi @NichD, thanks for the clarification, I think I have a better idea of your project's goal. It does seem that valis might not be suited to the 2nd part of this task, as registering the H&E stack to the fluorescent stack using valis would probably involve aligning corresponding Z-planes between the 2 modalities. It seems this could work if the fluorescent images were collected the same/similar regular intervals as the H&E slices though (e.g. every 4um). If that is the case, let me know and maybe I could put something together. If not, I'm afraid I don't have much experience with 3D registration, and haven't really encountered this challenge before :( Regarding the parameters I suggested, I should clarify that setting To create that figure, I used PyVista to create the 3D image. The code was something like this, where Hopefully some of that is useful, and if you have any more questions, please don't hesitate to ask :) Best, |
Beta Was this translation helpful? Give feedback.
-
|
@cdgatenbee thank you for that more detailed explanation. To respond to some of your comments, the fluorescent images were captured on the same tissue, but prior to fixation, making it a pretty challenging non-rigid registration task with considerable deformation expected. Regarding the setting In terms of volumetric cross-modal registration, have you explored any tools (open or closed source) that you've had success with? No worries if you haven't explored that. Thank you for pointing me toward PyVista! This looks like a great tool and I'm looking forward to exploring it. Much appreciated!! Incidentally, if you're developing a Docker image and build process for VALIS and you would like an additional tester, I'm happy to discuss this. I think, for the deploy that I'm using, a stable image would be preferred to having multiple environments on the shared computation resource I currently use. I'm happy to explore images built in Windows or Linux. Please let me know if I can be of assistance. Thank you again for your thoughts and suggestions. Much appreciated!! |
Beta Was this translation helpful? Give feedback.
-
|
Hi @NichD, Unfortunately, I don't have any experience with volumetric cross-modal registration, and so haven't explored any relevant tools :( It does sound like a very interesting task though, I just don't have any data to experiment with. That would be great if you could test the Docker image! I don't have easy access to a Windows machine, so it would be extremely helpful if you are able to test it on a PC. I'd share the Dockerfile now, but I switched to using poetry to install python packages so I could have a lockfile, and so the Dockerfile wouldn't work given the set of files currently on github. I should be pushing that update with the lockfile soon though, and will also put the Docker image on DockerHub. Best, |
Beta Was this translation helpful? Give feedback.
-
|
Hi @cdgatenbee - That additional detail regarding DeepFlow and displacements was helpful, thank you. I haven't noticed any disjoints in the transformed images using the default settings, but I'll pay attention to it. I'd be happy to test the Docker image and build in a Windows (and Linux) environment. I had originally attempted to use VALIS on my local Windows laptop, but ran into issues with the Maven - Scipy dependency chain. I think this is a great solution to try. Please let me know when those updates are ready and I'll check them out. Cheers, |
Beta Was this translation helpful? Give feedback.
-
|
Hi @NichD, Best. |
Beta Was this translation helpful? Give feedback.
Uh oh!
There was an error while loading. Please reload this page.
-
Hello! I've been using VALIS for a month now and found it a very useful and robust tool for registering WSI histology stacks. Thank you!!
This task is part of a larger workflow I'm exploring that will culminate in cross-modal registration of a volumetric histology dataset (registered histology stack mentioned above, effectively a volumetric dataset) with a live-tissue fluorescent dataset of the same sample prior to histological processing. This fluorescent dataset may be single plane, or volumetric.
I know from reading the docs that VALIS was designed with the intent of registering fluorescent and histology data. I wondered if this capability has been developed / explored with respect to volumetric data. Is VALIS limited to 2D registration problems, or is registration of volumetric data possible? If yes, can you provide any guidance on how that process is set up? If not, have you encountered in your travels any other image registration tools that might be well suited to this task?
Thanks in advance!!
Cheers,
Nicholas Dana
Beta Was this translation helpful? Give feedback.
All reactions